Hey, how’s your biotin? What? No it’s not an organic metal, maybe you call it B7? You’re probably fine, but have you been depressed, lethargic or losing your hair lately? Biotin is pretty important; it’s necessary for metabolism within our cells, so I make sure I never leave home without it. It’s rare for someone to have a biotin deficiency, but if you want to know your levels, give me a drop of your blood, and I’ll have an answer from you in 10 minutes. How? Oh just my self-powered integrated microfluidic blood analysis system (but I like to call it SIMBAS for short).

The SIMBAS emerged from Berkeley and was featured on the inside cover of the 2011 Issue 5 of Lab on a Chip. Luke Lee et al. describe a device capable of picomolar detection in “Stand-alone self-powered integrated microfluidic blood analysis system (SIMBAS).” The device can analyze whole blood without a lot of bells and whistles to get in the way. I’m not saying that this doesn’t have a clever design; it just doesn’t have a battery, moving parts or specialized readers. All you need is some blood and a microscope. The device has five independent analysis streams. There are three parts to a stream, the filter, detector and the suction chamber. All of these components are featured in a slab of PDMS sandwiched between two normal glass slides. Before I describe how the features physically work together, it’s important to note the device’s driving force: low pressure. Before use, the SIMBAS is placed in a low-pressure vacuum, degassing it through the single points of entry of each stream, creating a vacuum. When blood is placed at the inlet the vacuum slowly draws it through the system.
Filtration
After the vacuum sucks in the blood, the red and white blood cells must be filtered out. One major strength of this device is that it receives unadulterated blood. However, in order to prevent the cells from muddling the detection region of the device, they are filtered out by sedimentation. The floor of the 80 µm high, 50 µm wide sample channel opens up to a 2 mm wide, 2 mm deep circular well, which acts as a filter. The large, heavy cells sediment out and take up permanent residence at the bottom depending on the flow rate of the sample. The biomarkers of interest are much lighter and will continue through without joining the blood cells. This produces plasma that is 100% blood cell free, ready for detection. Platelets sediment at lower rates, so they will still remain in the plasma, but this doesn’t seem to be an issue.
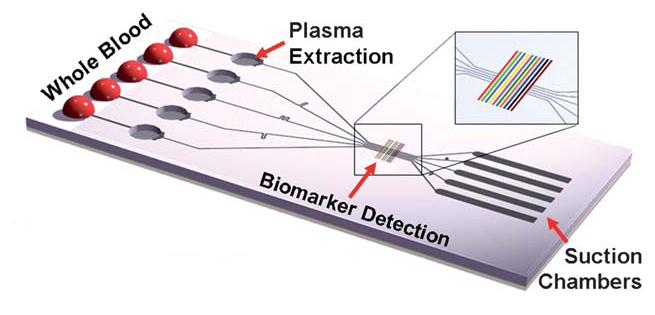
Detection
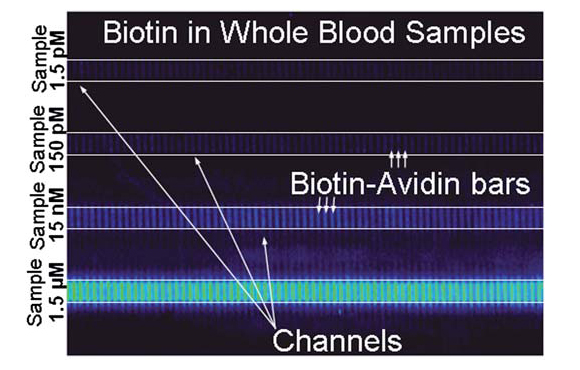
Now that the sample has been fully prepared, it is ready for detection. Remember that the PDMS was placed between two glass slides? The authors immobilized streptavidin bars on the underside of the glass roof. This leaves the streptavidin attached to the ceiling, letting it bind with the biotin that flows by. Streptavidin and biotin have one of the strongest known protein-ligand bonds, making it a perfect choice for this device. When tested, blood was spiked with fluorescently-labeled biotin at 1.5 pM. Many other detectors could be immobilized along the same stream or the other four streams. The five parallel streams allow a way to eliminate errors, or test for biomarkers that are not compatible with each other.
Volume Control
However, we’re still missing the last important feature, the suction trough, an empty region after the detectors. The volume of the trough determines how much sample is drawn through the system. Once the trough is full, flow stops, regardless of how much blood is ready to enter. Modifying the trough volume provides a method to increase the sensitivity of the device. A larger sample volume gives the SIMBAS a better chance to detect a biomarker at low concentrations.
Final Thoughts
The current setup only needs 5 µL of whole blood for each stream. To put that into perspective, modern glucometers need 1 µL at most to glucose levels. It is (relatively) a bit more blood, but you wouldn’t be doing this every day. After 10 minutes, biotin at 1.5 pM can be detected by removing the top slide and looking at it under a microscope. That’s a pretty low detection in my book. Crazy low. I’m sitting here trying to think of an accurate needle in a haystack analogy, but it’s not coming. Overall, this is a pretty innovative, yet simple device, and I’ll tell you what I think of its merits and things I’d like to see developed.
Merits:
- To start, this is has a great design for a point-of-care device, especially in a resource-poor setting. It has no external or moving parts, and requires no power. Sometimes in microfluidics, things can get very complicated with the number of channels, pumps, reagents etc., but this has a very clean and trouble-resistant design. It can be pre-packaged under low pressure so that a user only has to open it to activate the vacuum and use it within a couple of minutes.
- The fact that it can receive whole blood also makes it great for point-of-care. Some lab-on-a-chips actually depend on many sample preparation steps or external machines. But all the steps that are needed for it to do its job in isolation should be included, just like SIMBAS. The time between filtration and detection is pretty quick, which is important because the proteins found in the plasma change after longer separations.
- This doesn’t require much sample, and is still able to detect biotin at the low concentration of 1.5 pM. I’m not sure how clinically relevant that number is for biotin, but if you can detect that, you can play with the configuration to bring that down (or raise it much more easily).
What I’d like to see:
- The authors state that the biotin-streptavidin detection could be replaced by many other couples. I don’t doubt this, but I would like to see what detection levels they could achieve, since biotin-streptavidin has one of the strongest protein-ligand bonds.
- In the paper, the authors mention that the detection strips coated on the ceiling were applied on the same day the assay was run. I’d like to know what the shelf life of a prepared card would be, which could really impact their usefulness and value.
- As I mentioned previously, the authors detected fluorescent-tagged biotin in the blood, and examined this under a microscope. You can see the captured biotin fluorescing in their figure. But biotin isn’t naturally tagged with a fluorophore, which makes me wonder how they would normally detect a wild biomarker. Perhaps there is a noticeable difference under the microscope; otherwise they will need to introduce some tag in the blood that will act as a secondary binding agent.
- The last thing I’d like to see is the microscope. Well, I’d like to see it removed. This microfluidics system would be even greater if it was completely stand-alone. Daniel Fletcher (also from UC Berkeley) has developed a cell phone-powered field microscope capable of fluorescence microscopy, so it isn’t impossible in a resource-poor setting, but it’s just my wishful thinking.
References

Dimov, I., Basabe-Desmonts, L., Garcia-Cordero, J., Ross, B., Ricco, A., & Lee, L. (2011). Stand-alone self-powered integrated microfluidic blood analysis system (SIMBAS) Lab on a Chip, 11 (5) DOI: 10.1039/C0LC00403K